Tutorial¶
Creating Graphs¶
Let’s say that you wanted to create the following graph:
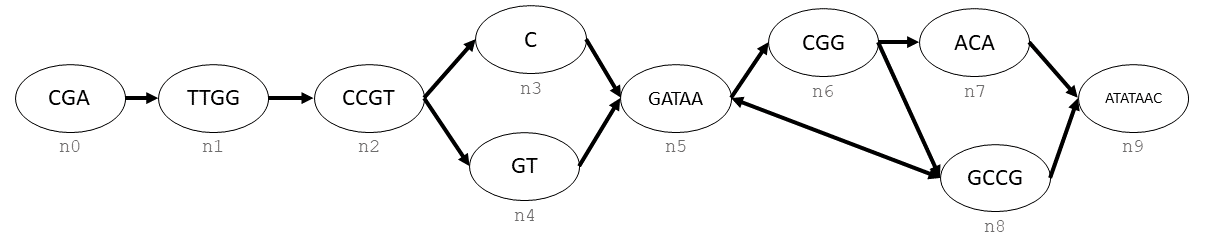
This graph is a combination of nodes (labelled as n0, n1, …, n9) and directed edges (arrows).
Graph Objects¶
Edges and nodes are accessed through an implementation of the bdsg.handlegraph.HandleGraph interface. Individual nodes in the graph are pointed at by bdsg.handlegraph.handle_t objects.
Paths exist in graphs that implement the bdsg.handlegraph.PathHandleGraph. Paths are accessed through bdsg.handlegraph.path_handle_t, which is a series of bdsg.handlegraph.step_handle_t linked together. Each bdsg.handlegraph.step_handle_t points to the node in that step, and also contains directional information regarding the nodes preceeding and following it.
Handles are pointers to specific pieces of the graph, and it is not possible to operate on them directly, aside from comparing whether the objects are equal. To get information regarding the object that each handle is pointing to, it is necessary to use the corresponding get accessor method on the graph that issued the handle.
Reference materials for these methods can be found at the Python API, as well as the Sorted Glossary of Methods, which contains lists sorted by object type for Accessor Methods, Mutator Methods, and Iteratator Methods.
Making a Graph¶
First, we must create the graph, then make each node and keep track of their handles. We’re going to be using the HashGraph, bdsg.bdsg.HashGraph, which is a good all-around graph that implements bdsg.handlegraph.MutablePathDeletableHandleGraph.
from bdsg.bdsg import HashGraph
gr = HashGraph()
seq = ["CGA", "TTGG", "CCGT", "C", "GT", "GATAA", "CGG", "ACA", "GCCG", "ATATAAC"]
n = []
for s in seq:
n.append(gr.create_handle(s))
Now we link together these nodes using their handles. Note that each of these handles is directional, and we create each edge from the first handle to the second. In order to create both of the edges between n5 and n8 (since each can follow the other) we use create_edge twice.
gr.create_edge(n[0], n[1])
gr.create_edge(n[1], n[2])
gr.create_edge(n[2], n[3])
gr.create_edge(n[2], n[4])
gr.create_edge(n[3], n[5])
gr.create_edge(n[5], n[6])
# Connect the end of n5 to the start of n8
gr.create_edge(n[5], n[8])
gr.create_edge(n[6], n[7])
gr.create_edge(n[6], n[8])
gr.create_edge(n[7], n[9])
gr.create_edge(n[8], n[9])
# Connect the end of n8 back around to the start of n5
gr.create_edge(n[8], n[5])
Traversing Edges¶
If we wanted to traverse these edges, we could do it using the iterator method bdsg.handlegraph.HandleGraph.follow_edges().
def next_node_list(handle):
lis = []
gr.follow_edges(handle, False, lambda y: lis.append(y))
return lis
print(f'n0: {gr.get_sequence(n[0])}')
next_node = next_node_list(n[0])[0]
print(f'n1: {gr.get_sequence(next_node)}')
next_node = next_node_list(next_node)[0]
print(f'n2: {gr.get_sequence(next_node)}')
Which will output the following:
n0: CGA
n1: TTGG
n2: CCGT
Creating a Path¶
Generating a linear sequence from this graph could be done in infinitely many ways, due to the interal loop between n5, n6, and n8. If we wanted to define a single consensus sequence, we would do this by defining a path.
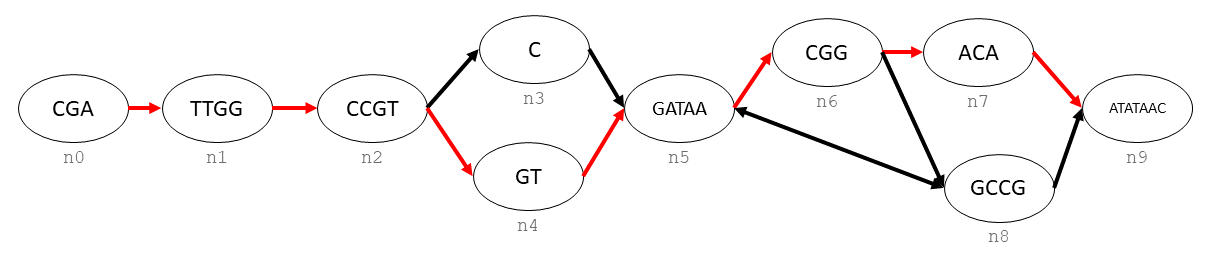
To create the hilighted path, we would need to create a bdsg.handlegraph.path_handle_t in the graph, and then append each bdsg.handlegraph.handle_t to the end of the path.
path = gr.create_path_handle("path")
gr.append_step(path, n[0])
gr.append_step(path, n[1])
gr.append_step(path, n[2])
gr.append_step(path, n[4])
gr.append_step(path, n[5])
gr.append_step(path, n[6])
gr.append_step(path, n[7])
gr.append_step(path, n[9])
Warning
bdsg.handlegraph.MutablePathHandleGraph.append_step() will not stop you from appending nodes that are not connected to the preceeding node.
# the following code runs without error
badpath = gr.create_path_handle("badpath")
gr.append_step(badpath, n[0])
gr.append_step(badpath, n[3])
Traversing a path¶
To traverse a path, we need to fetch a series of bdsg.handlegraph.step_handle_t from the graph. Note that although we are effectively asking the path for these items in it, all accessor methods are a part of the bdsg.handlegraph.PathHandleGraph object.
step = gr.path_begin(path)
while(gr.has_next_step(step)):
# get the node handle from the step handle
current_node_handle = gr.get_handle_of_step(step)
# ask the node handle for the sequence
print(gr.get_sequence(current_node_handle))
# progress to the next step
step = gr.get_next_step(step)
current_node_handle = gr.get_handle_of_step(step)
print(gr.get_sequence(current_node_handle))
Which will output the following:
CGA
TTGG
CCGT
GT
GATAA
CGG
ACA
ATATAAC
Saving and Loading Graphs¶
Graphs that implement bdsg.handlegraph.SerializableHandleGraph, such as bdsg.bdsg.HashGraph, can be saved and loaded through the bdsg.handlegraph.SerializableHandleGraph.serialize() and bdsg.handlegraph.SerializableHandleGraph.deserialize() methods.
Graph File Example¶
If you wish to save the graph from the above session, that can be done with:
gr.serialize("example_graph.hg")
This can be loaded into a new python session by using:
from bdsg.bdsg import HashGraph
gr = HashGraph()
gr.deserialize("example_graph.hg")
Loading in Pre-Existing Data¶
Each graph implementation knows how to read files in its respective file format.
For example, provided that data has been serialized in PackedGraph format, it is possible to read it directly from a file with bdsg.bdsg.PackedGraph. Download this graph and load it into python with:
from bdsg.bdsg import PackedGraph
brca2 = PackedGraph()
brca2.deserialize("cactus-brca2.pg")
We can poke around this data and get the sequence of the path with:
path_handle = []
handles = []
brca2.for_each_path_handle(lambda y: path_handle.append(y) or True)
brca2.for_each_step_in_path(path_handle[0],
lambda y: handles.append(brca2.get_handle_of_step(y)) or True)
sequence = ""
for handle in handles:
sequence += brca2.get_sequence(handle)
print(sequence[0:10])
print(len(sequence))
Note how we are using or True in the iteratee callback lambda functions to make sure they return True. If a callback returns False or None (which is what is returned when you don’t return anything), iteration will stop early and the for_each call will return False.
Reading in a Graph from vg¶
Graph assembies can be created with vg. Many .vg files that vg 1.28.0 or newer produces will be in HashGraph format, directly loadable by bdsg.bdsg.HashGraph.deserialize().
To check a file, you can use vg stats --format, like so:
vg stats --format graph.vg
If you see one of format: HashGraph or format: PackedGraph, you can probably (but not always; see the note on encapsulation) load the graph with bdsg.bdsg.HashGraph or bdsg.bdsg.PackedGraph, respectively.
However, in some circumstances, you will need to convert the graph to one of those formats. Some graphs (most notably, graphs from vg construct) will report format: VG-Protobuf, and .xg files will report format: XG. These graphs will need to be converted to HashGraph or PackedGraph format, and then loaded with the appropriate class. For example, you can do this:
vg convert --packed-out graph.vg > graph.pg
The resulting PackedGraph file can be loaded with bdsg.bdsg.PackedGraph.deserialize().
from bdsg.bdsg import PackedGraph
graph = PackedGraph()
graph.deserialize("graph.pg")
To use bdsg.bdsg.HashGraph instead, substitute --hash-out for --packed-out.
Older vg Graphs with Encapsulation¶
Versions of vg before 1.28.0 would encapsulate HashGraph and PackedGraph graphs in a file format that vg can read but libbdsg cannot. Consequently, some older graph files will be reported as format: HashGraph or format: PackedGraph by vg stats --format, but will still not be readable using libbdsg.
If you encounter one of these files, you can use vg view --extract-tag to remove the encapsulation and pull out the internal file which libbdsg can understand. For example, for a file that reports format: PackedGraph but is not loadable by libbdsg, you can do:
vg view graph.vg --extract-tag PackedGraph > graph.pg
This also works for HashGraph files, by replacing PackedGraph with HashGraph.
Running the file through vg convert with vg 1.28.0 or newer will also solve the problem, but could take longer.